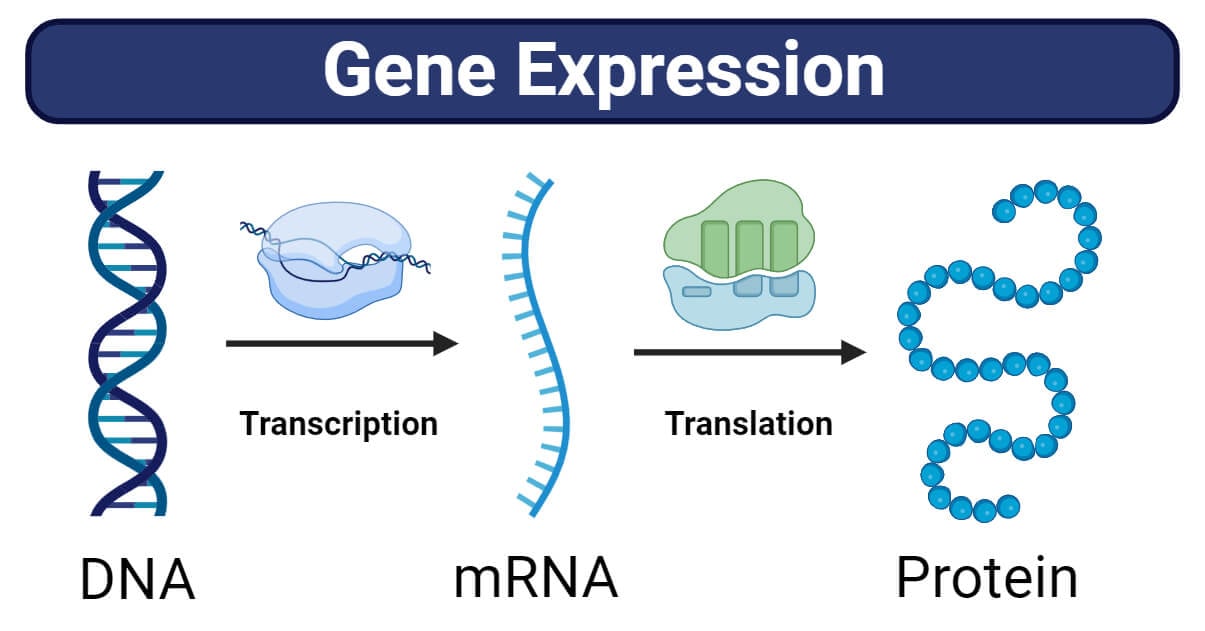
Protocol to use TopNet for gene regulatory network modeling using gene expression data from perturbation experiments - ScienceDirect
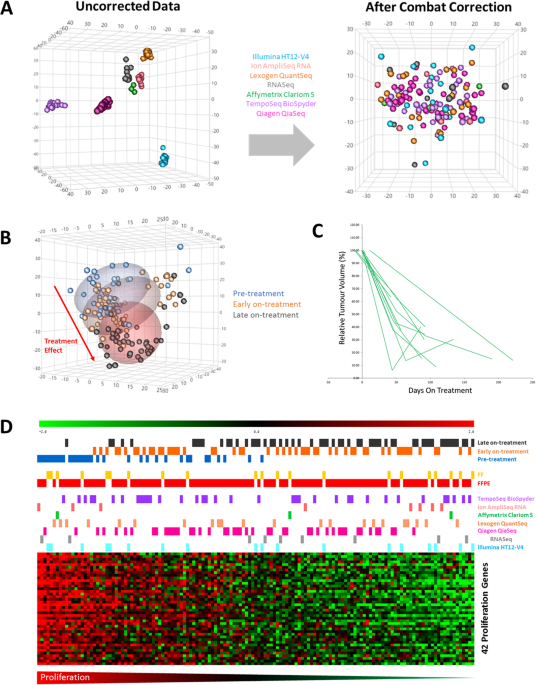
Unlocking the transcriptomic potential of formalin-fixed paraffin embedded clinical tissues: comparison of gene expression profiling approaches | BMC Bioinformatics | Full Text

Differential gene expression level measurement by microarray and qPCR.... | Download Scientific Diagram

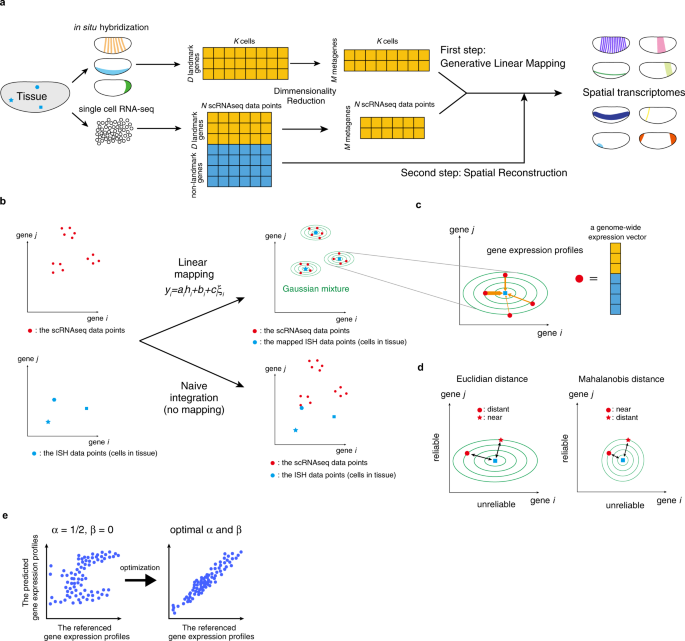
Using multiple measurements of tissue to estimate cell-type-specific gene expression via deconvolution | bioRxiv
![PDF] Measuring differential gene expression with RNA-seq: challenges and strategies for data analysis. | Semantic Scholar PDF] Measuring differential gene expression with RNA-seq: challenges and strategies for data analysis. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/75b8bf3156f8470555190d92662347f7506c3b29/4-Figure1-1.png)
PDF] Measuring differential gene expression with RNA-seq: challenges and strategies for data analysis. | Semantic Scholar
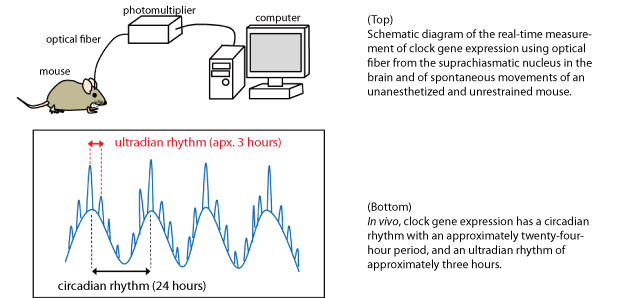
Successful Real-Time Measurement of Gene Expression in the Brain of Unrestrained Mice | Hokkaido University

Of Gene Expression and Cell Division Time: A Mathematical Framework for Advanced Differential Gene Expression and Data Analysis - ScienceDirect
![A novel approach for human whole transcriptome analysis based on absolute gene expression of microarray data [PeerJ] A novel approach for human whole transcriptome analysis based on absolute gene expression of microarray data [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2017/4133/1/fig-1-2x.jpg)
A novel approach for human whole transcriptome analysis based on absolute gene expression of microarray data [PeerJ]

ERROR MODELLED GENE EXPRESSION ANALYSIS (EMOGEA) PROVIDES A SUPERIOR OVERVIEW OF TIME COURSE RNA-SEQ MEASUREMENTS AND LOW COUNT GENE EXPRESSION | bioRxiv